The Integrative Genomics Building (IGB) seen above is home to the U.S. Department of Energy (DOE) Joint Genome Institute (JGI), a DOE Office of Science User Facility located at Lawrence Berkeley National Laboratory (Berkeley Lab), the DOE Systems Biology Knowledgebase (KBase), and the National Microbiome Data Collaborative (NMDC). Berkeley Lab Biosciences Area researchers are also co-located in the IGB.

The interactions between plants and the microbes in and around their roots influence plant health and development. Using controlled fabricated ecosystems or EcoFABs such as the one shown here help researchers conduct studies with reproducible parameters.
Director’s Perspective
Nigel Mouncey, Director, DOE Joint Genome Institute
In 2021, several researchers and staff supporting the JGI user community were recognized by multiple organizations.
Science Highlights

Mingqin “Mike” Shao in the plant room at the IGB checking one of the JGI flagship genome species, Brachypodium distachyon.
 |
Shattering Expectations: Novel Seed Dispersal Gene Found in Green Millet
Researchers at the Danforth Center, the JGI and the HudsonAlpha Institute for Biotechnology released a very high-quality reference green millet genome sequence and identified a gene related to seed dispersal in wild populations. |
 |
The Case for Conservation
The genome assemblies of high-quality reference sequences for the male and female fire moss plants (Ceratodon purpureus) are a testament to the multiple advances in sequencing technologies applied to the effort. |
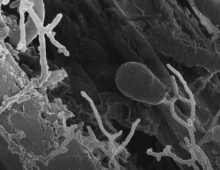 |
Gut Fungi: Unexpected Source of Novel Chemicals
Combing through the genomes of four anaerobic fungal species has revealed, for the first time, that this group is unexpectedly powerful: they can whip up dozens of complex natural products, including new ones. |
 |
Bacteria and Fungi Divvy Up the Work in Forest Floor
By analyzing the enzymes present in the complex organic matrix of the forest floor, researchers found that fungi are more active in degrading plant matter, and bacteria are more active in fixing and metabolizing nitrogen. |
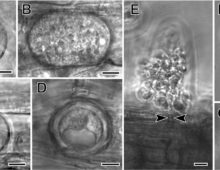 |
Olpidium, The Key to the Origin of Terrestrial Fungi
Taking advantage of the rapid development of binning methods, and with much more extensive taxonomic and genetic sampling, researchers confirmed the affinity between Olpidium and the non-flagellated terrestrial fungi. |
 |
Marine Microbe Contains Multitudes
In the ocean’s North Pacific Subtropical Gyre, microbes tend to stay localized at different depths. Scientists looked into how the bacterium SAR324 can be found throughout the water column. |
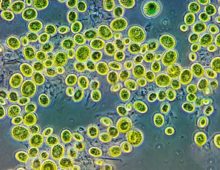 |
A One-Stop Shop for Analyzing Algal Genomes
PhycoCosm’s interactive browser allows researchers to look deep into more than 100 algal genomes. The genome portal reinforces the JGI’s new strategic focus on exploring algal biology, diversity, and ecology. |
 |
Climate Change Threatens Base of Polar Oceans’ Bountiful Food Webs
An international research team reported warm-adapted microbes edging polewards and may be displacing resident tiny algae much more easily than previously suspected. The trend could destabilize the delicate marine food web. |
 |
Refining the Process of Identifying Algae Biotechnology Candidates
JGI, LANL and NREL researchers combined expertise to screen, characterize, sequence and then analyze the genomes and multi-omics datasets for algae that can be used for large-scale production of biofuels and bioproducts. |
 |
Green Algae Reveal One mRNA Encodes Many Proteins
Gene expression in eukaryotes was long held to be monocistronic; a single gene makes messenger RNA, which encodes a single protein. Researchers have found numerous examples of polycistronic expression in green algae. |
 |
An Automated Tool for Assessing Virus Data Quality
A command-line tool called CheckV can be broadly utilized to gauge virus data quality. It helps researchers to follow best practices and guidelines for providing the minimum amount of information for an uncultivated virus genome. |
 |
A Natural Mechanism Can Turbocharge Viral Evolution
Researchers discovered that “diversity generating retroelements” (DGRs) in viruses are not only widespread, but also surprisingly active. They appear to generate diversity quickly, allowing these viruses to target new microbial prey. |
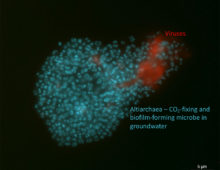 |
Plotting a Model for Virus-Host Warfare Deep Below Ground
Altiarchaea counter multiple attempts at virus infections because their genomes include sequences that code for CRISPR systems. These help bacteria resist foreign genetic elements by incorporating fragments from infecting viruses and phages. |
 |
An Age of CRAGE: Advances in Rapidly Engineering Non-model Bacteria
The JGI has demonstrated CRAGE as a versatile engineering system that allows scientists to conduct genome-wide screens and explore biosynthetic pathways. Now CRAGE is being applied to other synthetic biology problems. |
 |
Designer DNA: JGI Helps Users Blaze New Biosynthetic Pathways
A special issue of the journal Synthetic Biology puts the spotlight on JGI’s DNA design and synthesis superpower. In a series of case studies, scientific users share what they’ve discovered through their collaborations. |
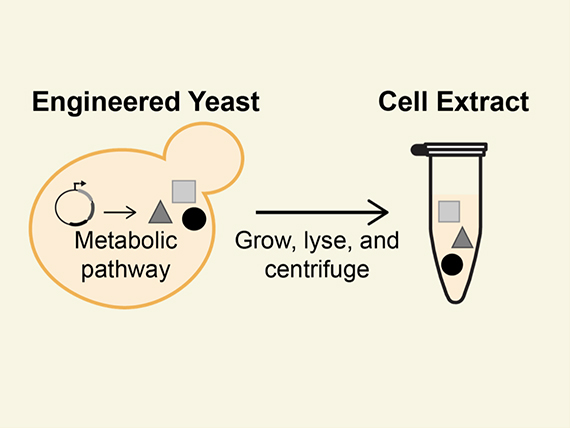 |
Boosting Small Molecule Production in Super “Soup”
In work enabled by the JGI’s Emerging Technologies Opportunity Program (ETOP), The University of Texas at Austin and Northwestern University researchers show yeast metabolic pathways work outside the cell environment. |
Impact: By the Numbers

This image shows Diane Bauer monitors a liquid handler processing sequencing samples. The JGI generated a record 467 Terabases of sequence in FY2021.
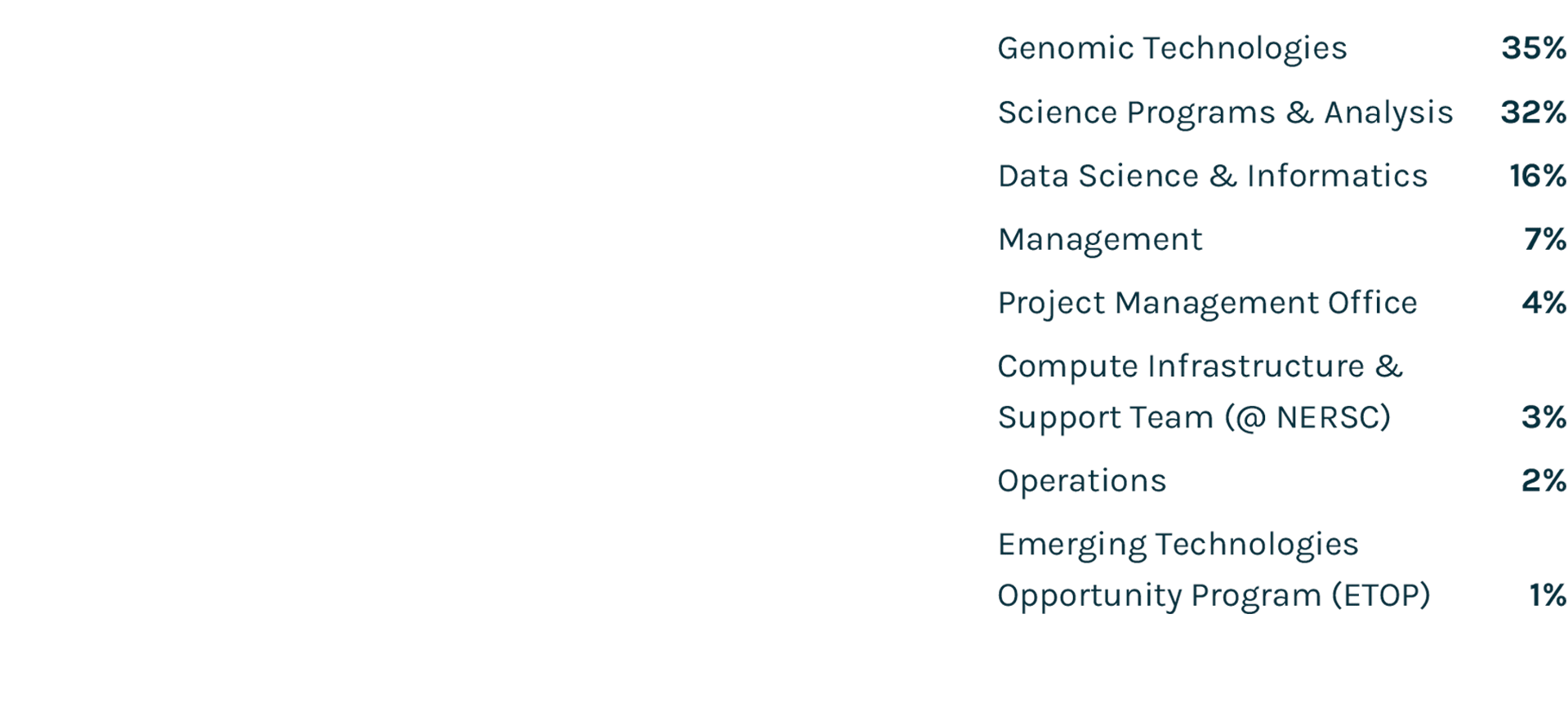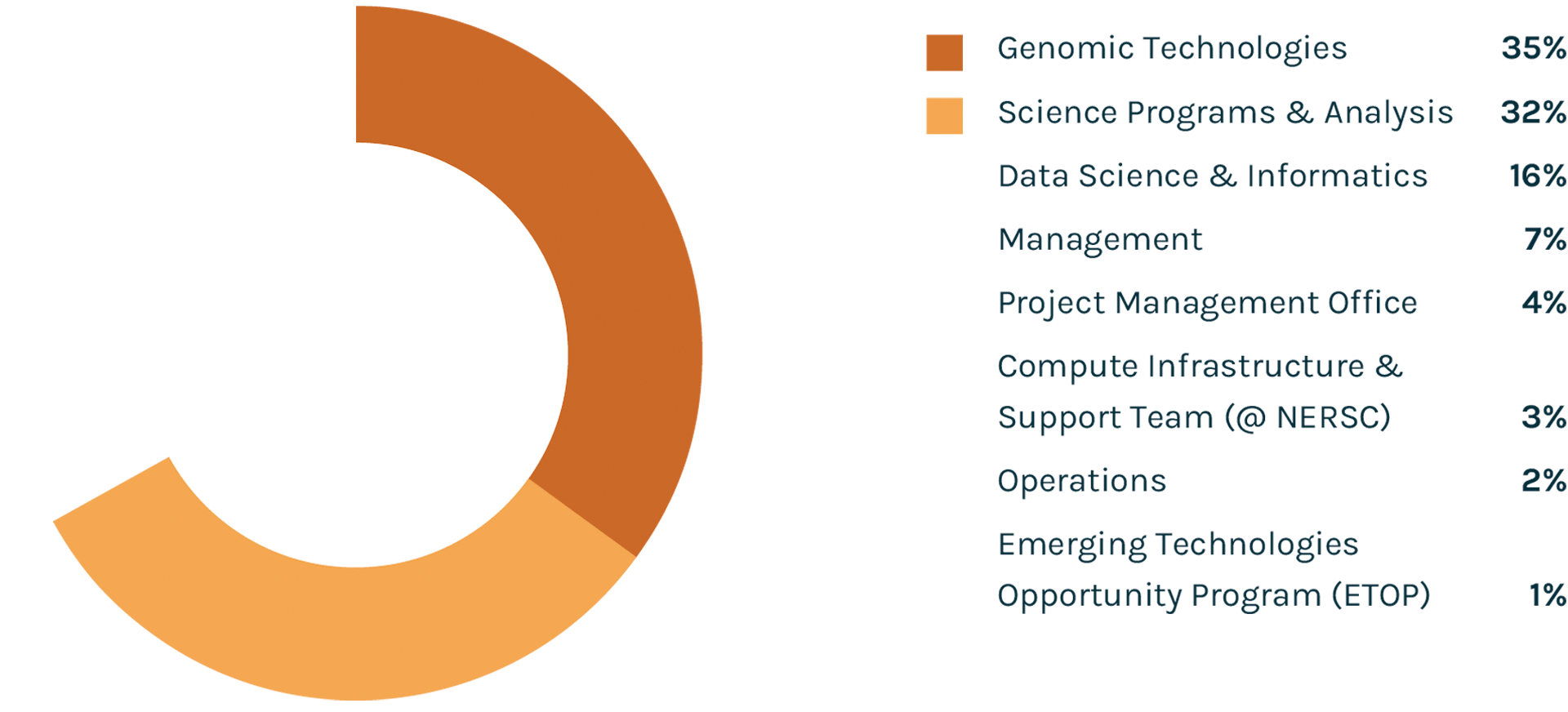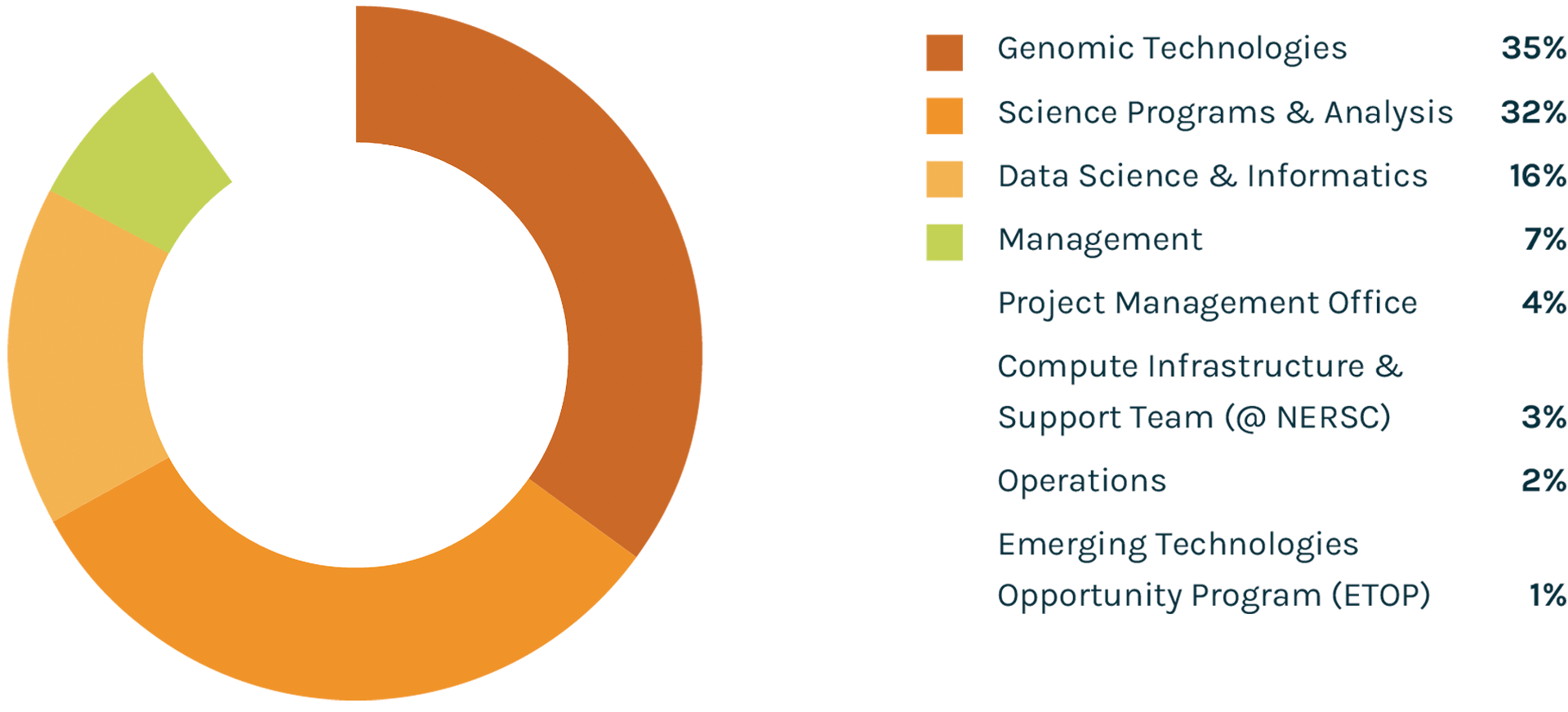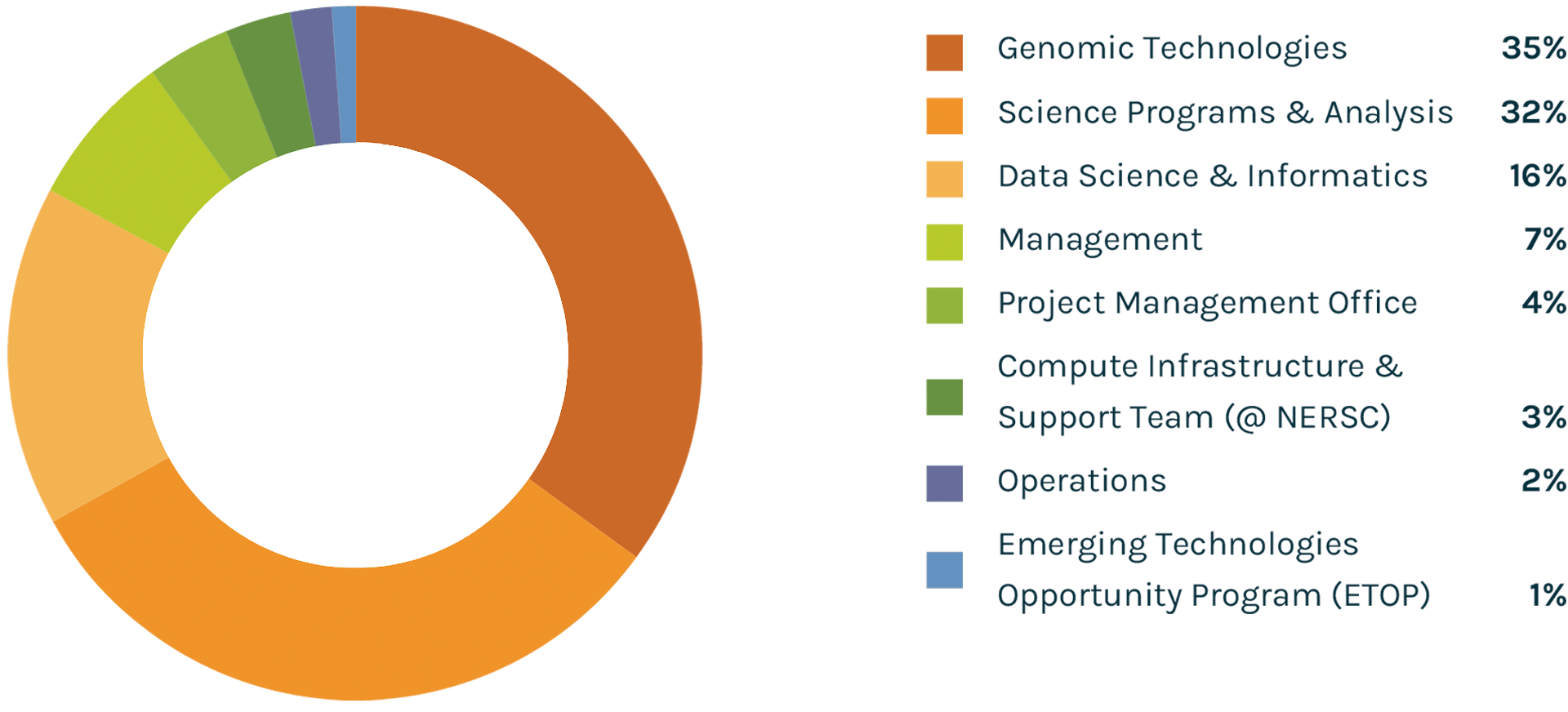
Spending Profile FY2021
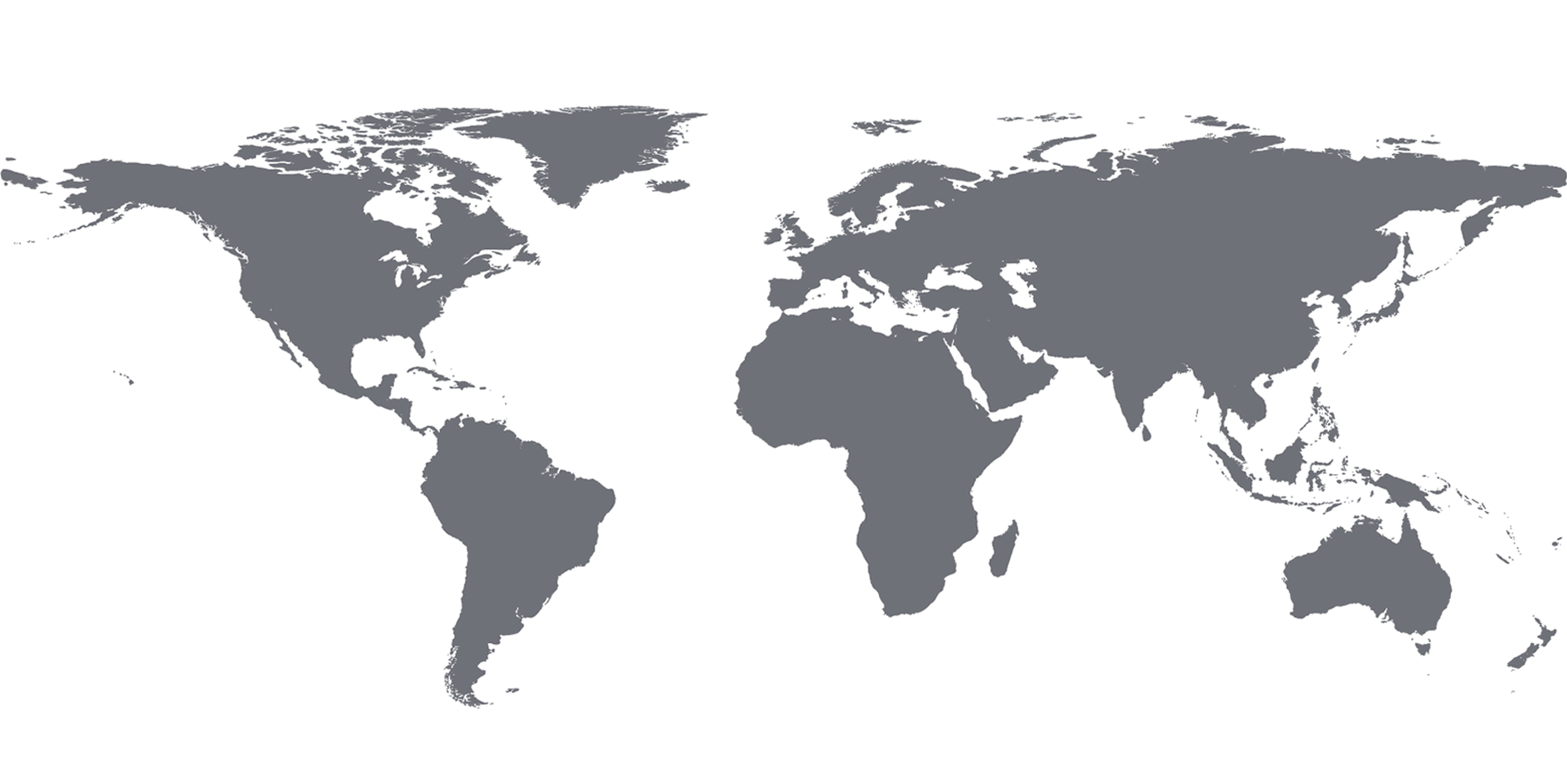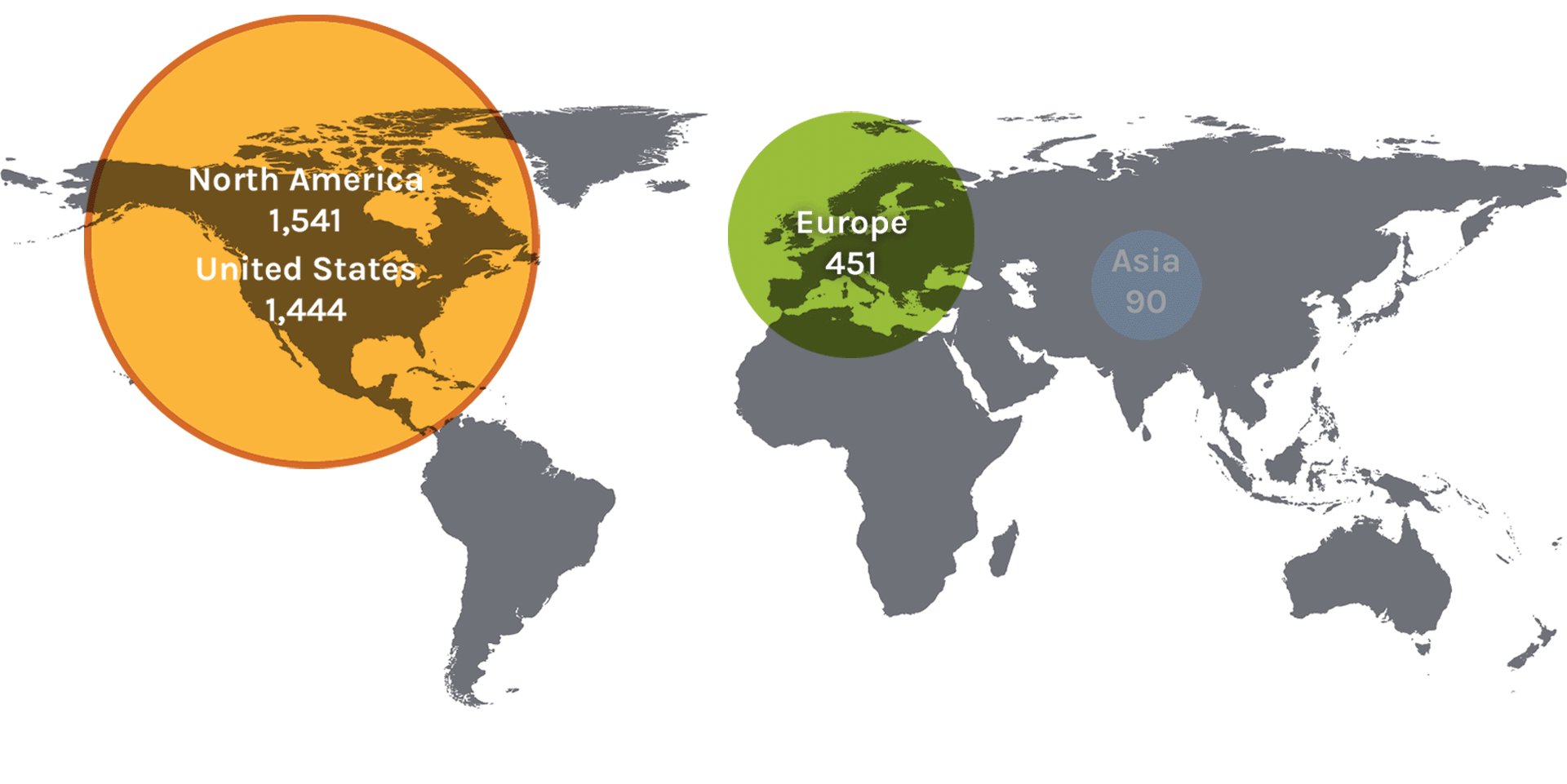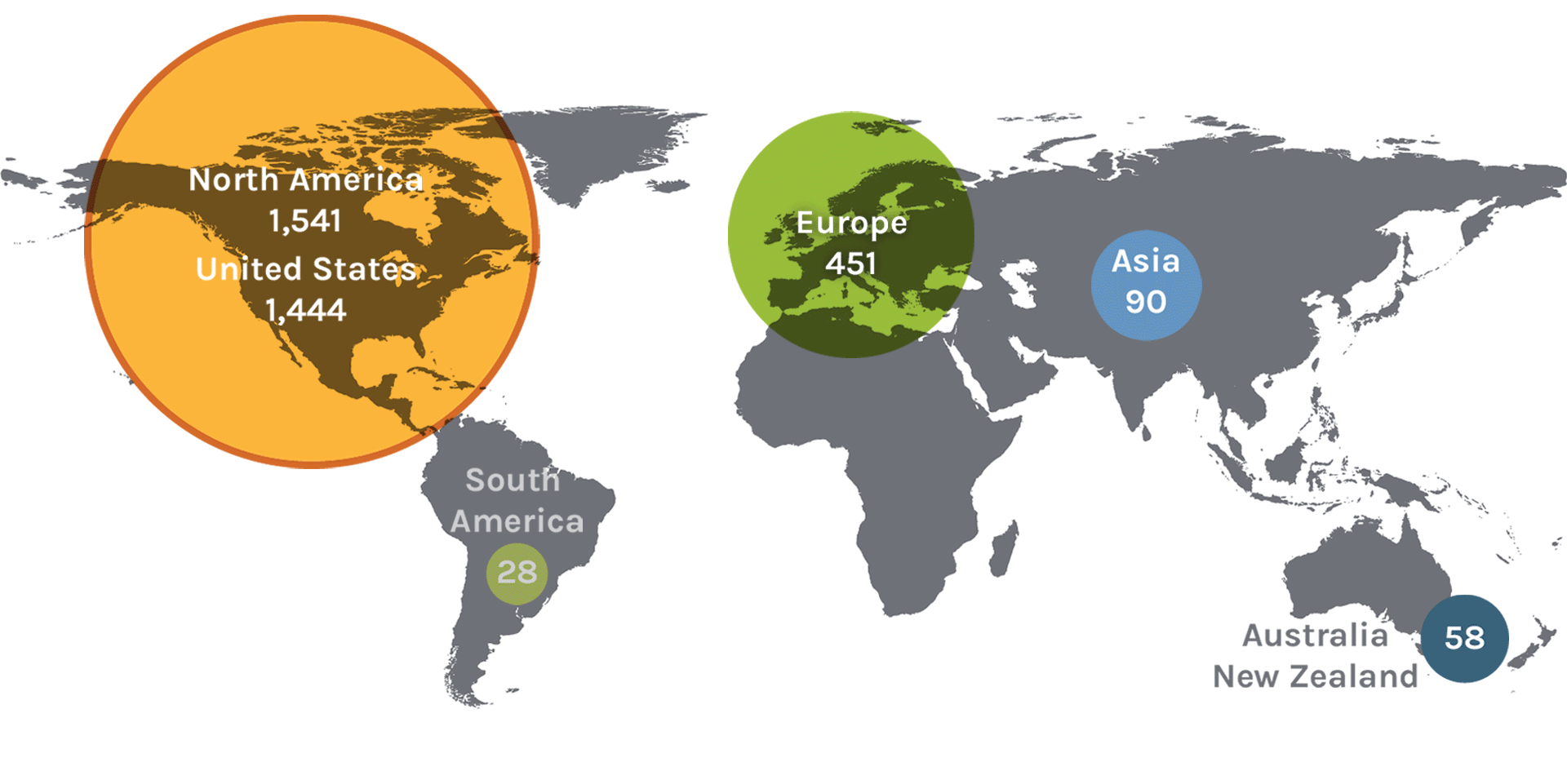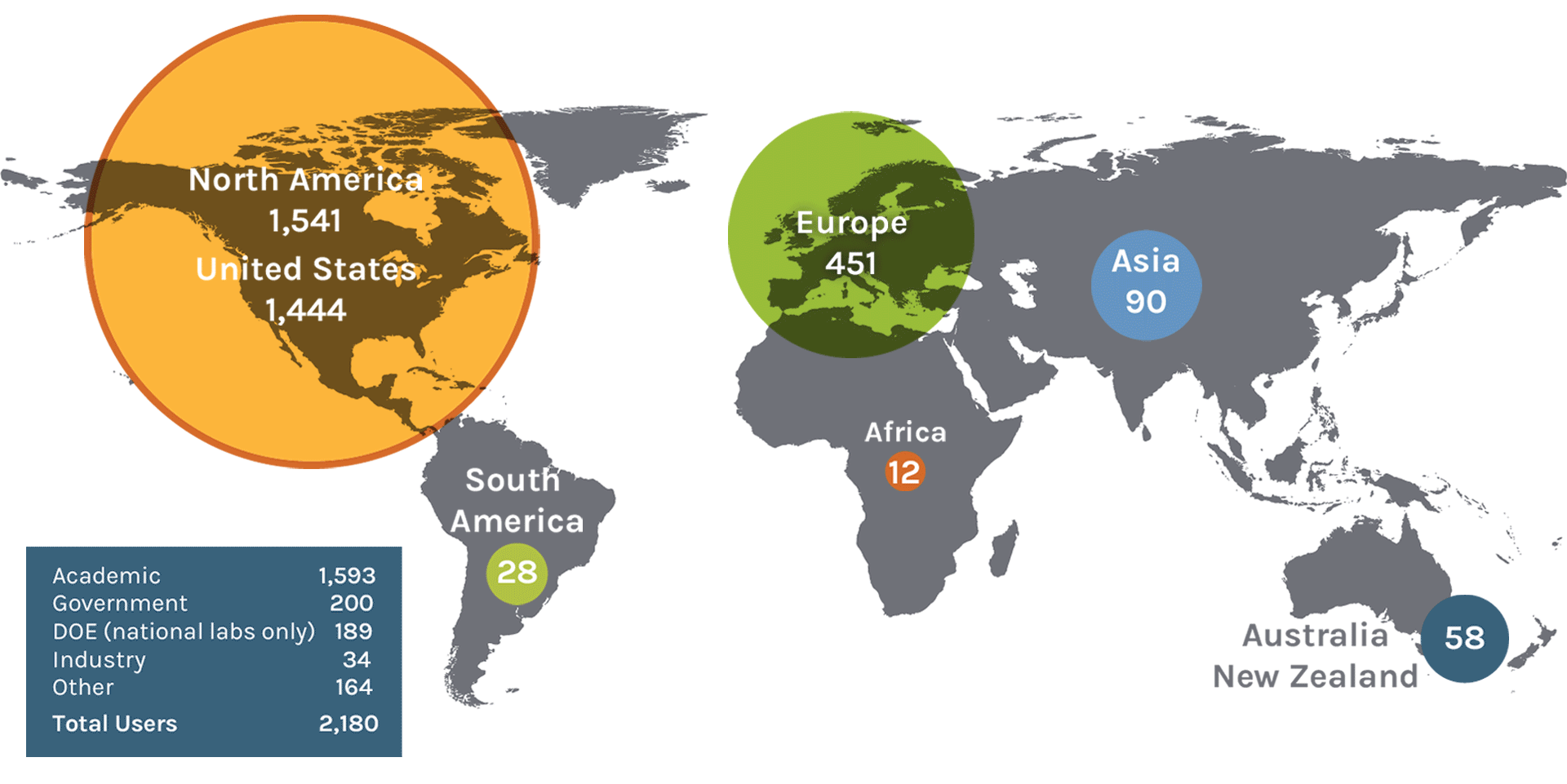
Users on the Map: 2,180
| North America | 1,541 | Denmark | 11 | Slovenia | 2 | Japan | 19 |
| United States | 1,444 | Estonia | 2 | Spain | 40 | Malaysia | 1 |
| Canada | 92 | Finland | 13 | Sweden | 19 | Oman | 1 |
| Mexico | 5 | France | 55 | Switzerland | 11 | Singapore | 3 |
| Germany | 95 | United Kingdom | 56 | South Korea | 5 | ||
| South America | 28 | Greece | 3 | Taiwan | 2 | ||
| Argentina | 1 | Hungary | 11 | Africa | 12 | Vietnam | 1 |
| Brazil | 20 | Iceland | 1 | Morocco | 2 | ||
| Chile | 1 | Ireland | 3 | Nigeria | 1 | Australia & New Zealand | 58 |
| Colombia | 2 | Italy | 31 | South Africa | 8 | Australia | 47 |
| Uruguay | 4 | Netherlands | 26 | Tunisia | 1 | New Zealand | 11 |
| Norway | 22 | ||||||
| Europe | 451 | Poland | 3 | Asia | 90 | ||
| Austria | 11 | Portugal | 8 | China | 36 | ||
| Belgium | 16 | Russia | 5 | India | 13 | ||
| Czech Republic | 5 | Serbia | 2 | Israel | 9 |
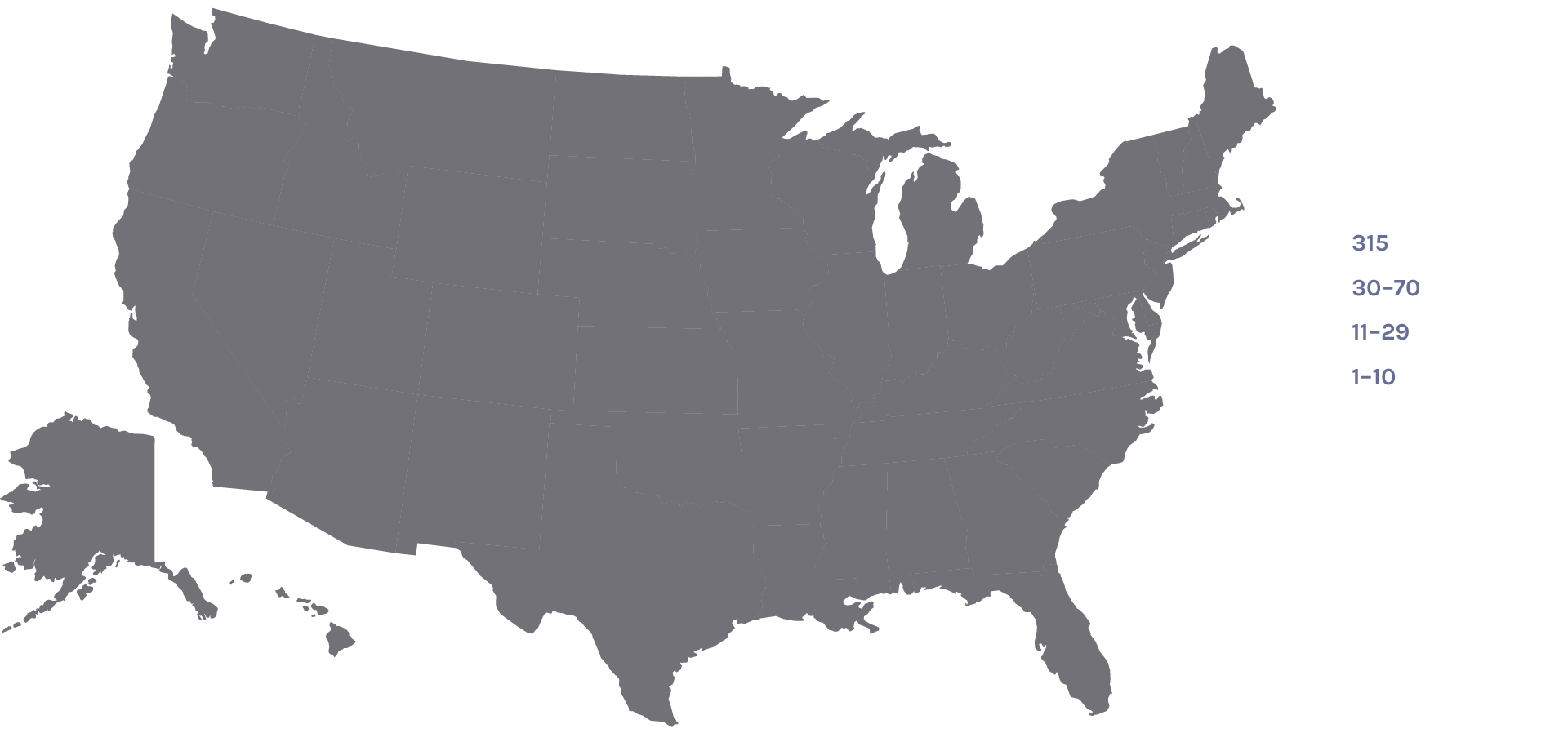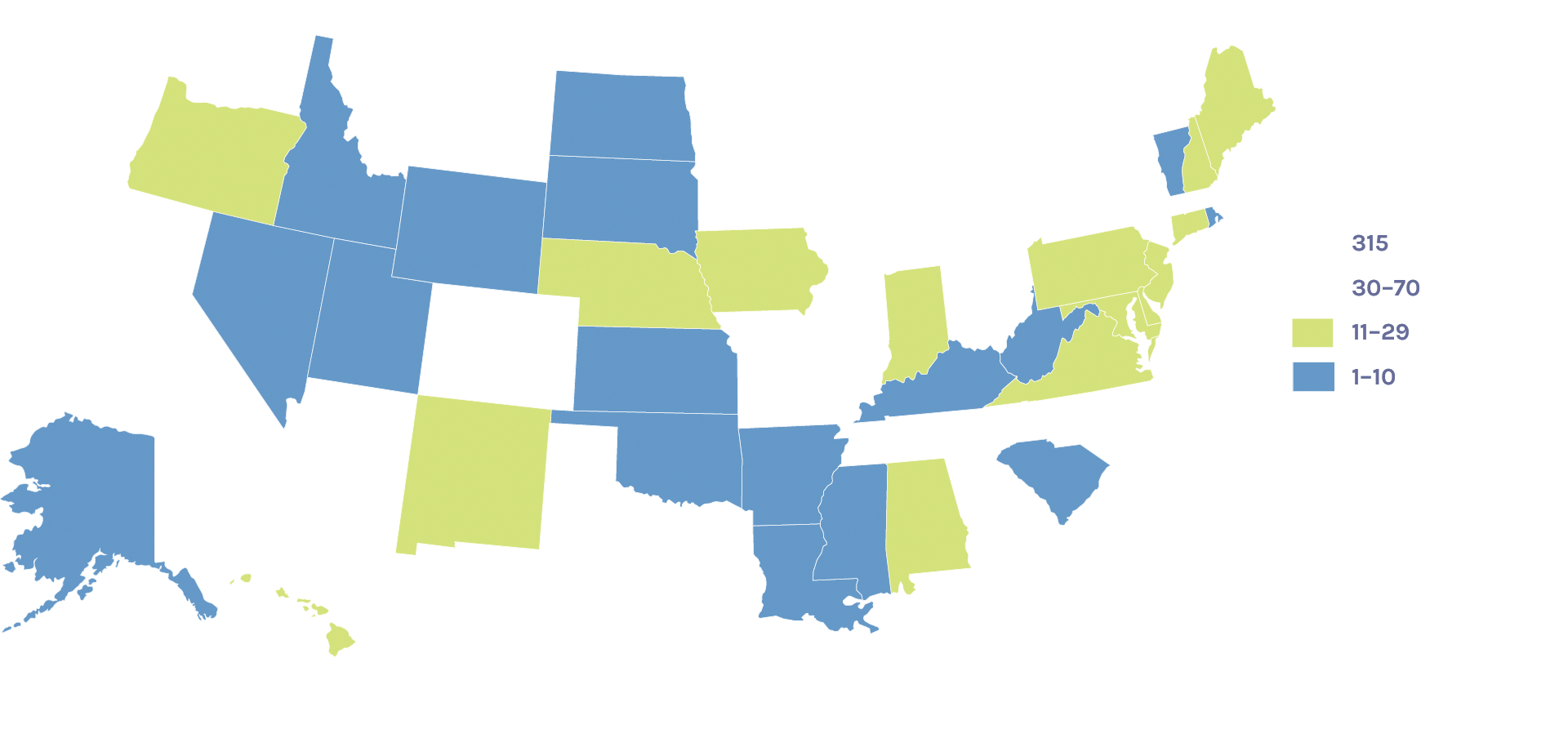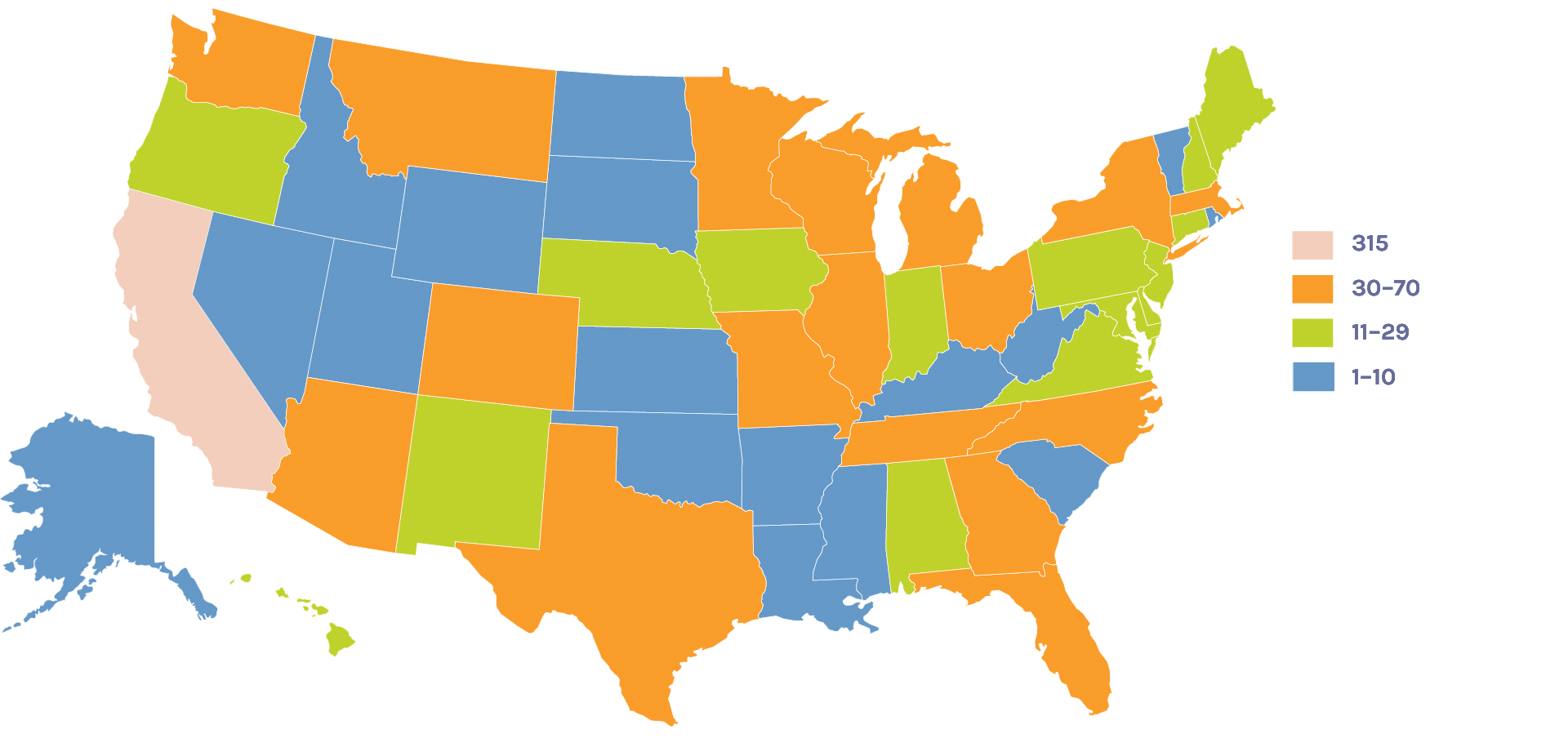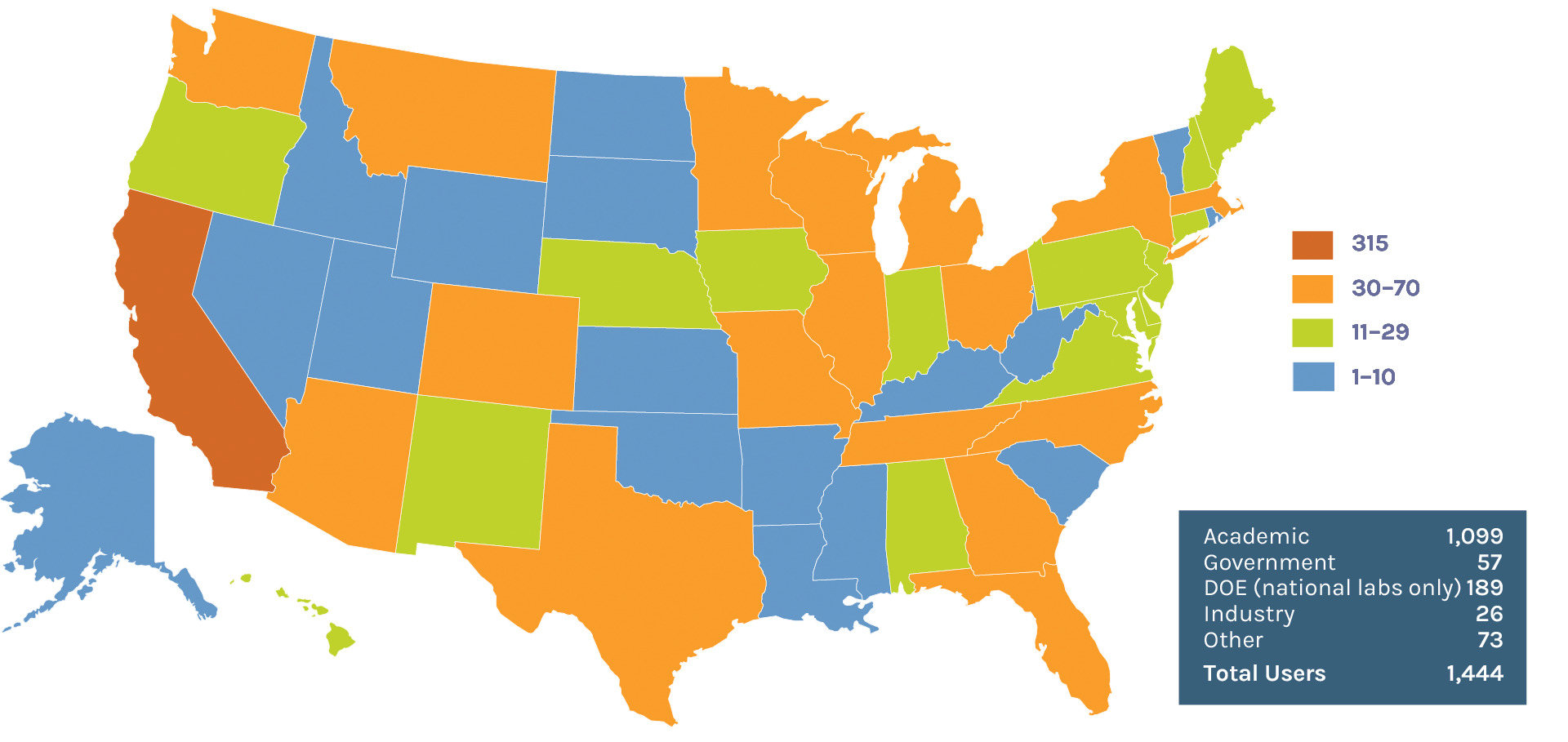
Users on the U.S. Map: 1,444
Cumulative Number of Projects Completed |
Cumulative Number of Scientific Publications |
 |
|
Sequence Output
(in billions of bases or GB)
The JGI supports short- and long-read sequencers, where a read refers to a sequence of DNA bases. Short-read sequencers produce billions of paired-end 150 basepair reads used for quantification, such as in gene expression analysis. Long-read sequencers currently average 60,000–70,000 bp reads and are used for de novo genome assembly. Combined short-read and long-read totals per year give JGI’s annual sequence output. The total sequence output in 2021 was 467,195 GB.
Sequencing Productivity
Billions of Base Pairs
User Letters of Intent/Proposals Submitted & Approved
Computational Infrastructure

The JGI has generated petabytes of high-quality sequence data and analysis; rapid and smooth access to the public datasets by the research community is enabled by high-performance computing resources and infrastructure. In this image, NERSC engineer James Botts studies a computer system cluster.
JGI Archive and Metadata Organizer (JAMO)11,010 million file records JAMO Archived Data Footprint11.683 Petabytes (PB) Data Downloads in FY214.201 million files; 1.646 PB |
Users of JGI Tools & DataThe Genome Portal provides unified access to all JGI genomic databases and analytical tools. A user can search, download and explore multiple data sets available for all JGI sequencing projects including their status, assemblies, and annotations of sequenced genomes. Launched in FY2021, the Data Portal allows JGI users to more easily access public data sets through a common set of metadata across the files that are submitted by each scientific program. The Genome Portal will be retired once the same features are available on Data Portal. |
Photography and cinemagraphs by Thor Swift, Berkeley Lab. Design by Creative Services, IT Division, Berkeley Lab.