JGI at 25: Mapping Switchgrass Traits with Common Gardens
Long-term research investments in switchgrass genomics are helping these grasses become a key player in biomass-based fuels. [Read More]
 Long-term research investments in switchgrass genomics are helping these grasses become a key player in biomass-based fuels. [Read More]
Long-term research investments in switchgrass genomics are helping these grasses become a key player in biomass-based fuels. [Read More] To make fuels and chemicals from plants, we’ll need new ways of processing bark, shoots and leaves. Hear from JGI User Michelle O’Malley about the gut fungi that could be the key to better biomass breakdown. [Read More]
To make fuels and chemicals from plants, we’ll need new ways of processing bark, shoots and leaves. Hear from JGI User Michelle O’Malley about the gut fungi that could be the key to better biomass breakdown. [Read More] Hear the 2022 cohort of UC Merced undergraduate and graduate students share how their summer internship experiences at the JGI have influenced their careers in science. [Read More]
Hear the 2022 cohort of UC Merced undergraduate and graduate students share how their summer internship experiences at the JGI have influenced their careers in science. [Read More]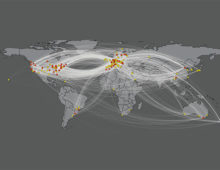 Researchers are forming communities to generate high quality genomes in partnership with the JGI to benefit scientists around the world. [Read More]
Researchers are forming communities to generate high quality genomes in partnership with the JGI to benefit scientists around the world. [Read More] Since 2016, the JGI has leveraged their work as pioneers in metagenomics to boost understanding of environmental viruses. At the time, the JGI expanded its existing Integrated Microbial Genomes & Microbiomes database to include a dedicated section for viruses; the latest release of IMG/VR currently features over 15 million viral genomes. [Read More]
Since 2016, the JGI has leveraged their work as pioneers in metagenomics to boost understanding of environmental viruses. At the time, the JGI expanded its existing Integrated Microbial Genomes & Microbiomes database to include a dedicated section for viruses; the latest release of IMG/VR currently features over 15 million viral genomes. [Read More] The dataset from a 2011 paper identifying microbial genes in cow rumen is now used for a hands-on undergraduate research course at four universities. [Read More]
The dataset from a 2011 paper identifying microbial genes in cow rumen is now used for a hands-on undergraduate research course at four universities. [Read More] Hands-on experience can be life changing for students. Hear former JGI interns share how these experiences have influenced their career journeys. [Read More]
Hands-on experience can be life changing for students. Hear former JGI interns share how these experiences have influenced their career journeys. [Read More] Every year, the JGI sequences around 35,000 samples — from plants, algae, bacteria, archaea, fungi, viruses — to support scientists around the world. Most of those researchers send their samples in from afar, without ever hearing much about the sequencing lab. So today, Chris Daum walks through the JGI’s sequencing pipeline, where there are freezers with names — but not doors — and robots handle a bunch of benchwork.
[Read More]
Every year, the JGI sequences around 35,000 samples — from plants, algae, bacteria, archaea, fungi, viruses — to support scientists around the world. Most of those researchers send their samples in from afar, without ever hearing much about the sequencing lab. So today, Chris Daum walks through the JGI’s sequencing pipeline, where there are freezers with names — but not doors — and robots handle a bunch of benchwork.
[Read More] In 2018, the JGI helped assemble the Hungate1000 catalog. To date it is the single largest effort to provide a cataloged and curated culture and genome sequence resource of rumen microorganisms. [Read More]
In 2018, the JGI helped assemble the Hungate1000 catalog. To date it is the single largest effort to provide a cataloged and curated culture and genome sequence resource of rumen microorganisms. [Read More] As part of the JGI’s 25th anniversary celebration, the 2022 Annual Meeting featured speakers whose talks shed light on how the JGI was established, all that it has contributed, and what they’re excited about in JGI’s future. [Read More]
As part of the JGI’s 25th anniversary celebration, the 2022 Annual Meeting featured speakers whose talks shed light on how the JGI was established, all that it has contributed, and what they’re excited about in JGI’s future. [Read More]