Found in a diverse range of environments, cyanobacteria play a significant role in the global atmospheric carbon cycle; in the oceans, they are responsible for reducing the carbon dioxide concentration in the atmosphere by as much as 400 parts per million (ppm). Researchers are interested in learning more about how cyanobacteria react to environmental stressors such as changes in light, temperature and nutrient availability.
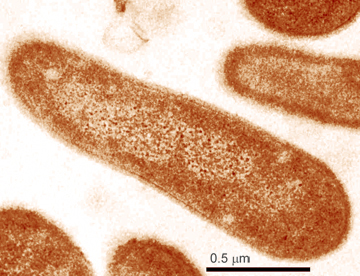
Photo: L.A. Sherman and D.M. Sherman, Purdue University
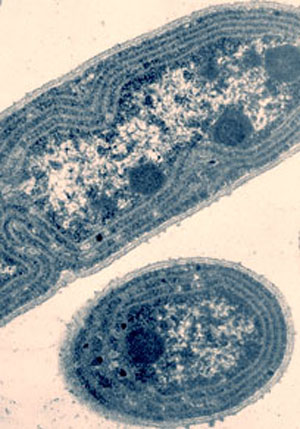
Photo: L.A. Sherman and D.M. Sherman, Purdue University
This project focuses on the cyanobacteria Synechococcus sp. PCC 7942 and Synechocystis sp PCC 6803, both among the first bacterial genomes to be sequenced. Both are model organisms for studying photosynthesis, carbon and nitrogen assimilation, hydrogen production and adaptation to environmental fluctuations and stresses. To learn more about these cyanobacteria, and how they react to stress, the researchers intend to focus on the transcriptome – the region of the genome that contains the instructions for building and maintaining cells by transcribing DNA into RNA – of these bacteria after growing the bacteria under a variety of conditions. By comparing the transcriptomic information of the two cyanobacteria, researchers will increase their understanding of how the bacteria can adapt their physiology and metabolism in response to various conditions and the resulting impact on the global carbon balance. The information will also help researchers improve the functional annotation of their genomes.
Principal Investigators: Konstantinos Mavrommatis, DOE Joint Genome Institute
Program: CSP 2010