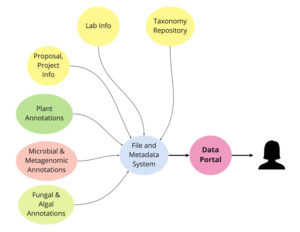
With a common set of “baseline metadata,” JGI users can more easily access public data sets. (Steve Wilson)
When the JGI Data Portal (https://data.jgi.doe.gov/) launched last year, it was only accessible through the plant portal Phytozome. Now the portal offers a way for users to more easily access public data sets through a common set of metadata.
For Steve Wilson, JGI’s Systems Engineering group lead, the Data Portal reflects a structural shift in data access. Users were previously limited to accessing data sets within groups, such as plants (via Phytozome) or fungi (MycoCosm). “We have made a concerted effort to create a common set of ‘baseline metadata’ across the files that are submitted by each scientific program,” he said. “If each kingdom submits the same category of data (key) for their files as a baseline, we can allow a user to collect all of the ‘protein FASTA files’ more easily.” He also elaborated on the following topics.
Q(uestion): What is the Data Portal’s scope?
A(nswer): The JGI Data Portal currently allows users to find files by searching file metadata (info describing the files)
We are currently limited to:
-
- Public data: datasets associated with completed projects that are eligible for public release, have completed their embargos, and numerous other requirements.
- Data that passes through a kingdom portal (assemblies + annotations)
- For IMG & Mycocosm: Data that is associated with an ITS project ID (AP or SP)
Q: What JGI Data Policy considerations do users accessing need to be mindful of?
A: The JGI Data Portal currently only presents public data (both restricted and unrestricted).
The Data Portal presents the users with the standard JGI Data Release Policy information when they request a download. When we have a calculation for automatically determining which datasets are unrestricted and which are not, we will be able to display that on JDP and allow users to filter on that parameter.
The current Data Restriction Policy requires that users know about the FY Funding Year, and the publication status in addition to the public/private status.
Q: Does Data Portal work well with KBase and NMDC?
A: We have reached out to KBase regarding use of our search API. They have expressed an interest in using this to find files based on file metadata criteria.
Wilson said that the Data Portal and Genome Portal will continue to run in parallel for now. Eventually, he added, the Genome Portal will be retired once the same features are available on Data Portal.