Microbes disprove long-held assumption that all organisms share a common vocabulary.
The Science:
Through single-cell genomics and metagenomics, researchers exploring the planet’s microbial diversity have found that not all organisms interpret a series of short genetic sequences to mean the same thing.
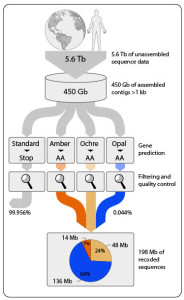
Workflow (depicted in the amount of DNA sequence data generated in trillion of nucleotide bases [Tb] and billion bases [Gb]) to identify the set of overlapping DNA segments that contain stop codon (Amber, Ochre, Opal) reassignments. (Patrick Schwientek)
The Impact:
The ability to study microbes in the wild helps researchers realize that some preconceived notions of how microbes behave may be erroneously based on data from the small fraction of microbes that have been cultivated and studied.
Summary
Four letters – A, C, G and T – make up the bases in DNA, and these same four letters repeating in a particular order genetically define an organism. Within this sequence are shorter, three-letter codes aptly named codons that represent amino acids, the building blocks of proteins that carry out a myriad of functions. There are 64 of these codons and nearly all of them code for the 20 amino acids. Three triplets are known as stop codons and used to signal the end of producing a protein. However, a recent study from the U.S. Department of Energy Joint Genome Institute shows that for some organisms, the instructions for these three codons mean anything but stop.
As reported in the May 23, 2014 issue of Science, the team led by DOE JGI Director Eddy Rubin focused on uncultivated microbes whose genomes had been described through single-cell genomics and metagenomics, and on a collection of viral sequences. Nearly six terabases of sequence data was analyzed from 1,776 samples collected from the human body and several sites around the world. “In this project, using metagenomics and single-cell genomics to explore uncultured microbes, we really had the opportunity to see how the genetic code operates in the wild,” Rubin said. “It is helping us get an unbiased view of how nature operates and how microbes manage our planet.”
Given that all organisms have genomes built on the same four letters, scientists had long assumed that they also all shared the same vocabulary and the 64 codons would be interpreted the same way across the board. What they found when they looked at uncultivated microbial sequences showed them that codons get reassigned more often that had been previously thought.
“Reassignment of all three stop codons was found but with different preferences by domain and habitat,” the team reported. “We observed distinct patterns of stop codon reassignment in the three domains of life, with bacteria showing only opal reassignments, ochre reassignments restricted to eukaryotes, and archaea devoid of codon reassignments. Among [DNA] viruses, we found both amber and opal reassignments.”
This work builds on a previous study in which DOE JGI researchers successfully employed single-cell genomics to shed insight on a plethora of microbes representing 29 “mostly uncharted” branches on the tree of life.
Contact
Eddy Rubin
DOE Joint Genome Institute
emrubin@lbl.gov
Funding
- U.S. Department of Energy Office of Science
Publication
Ivanova N. Stop codon reassignments in the wild. Science. 2014 May 23. 344(6186):909-913. doi: 10.1126/science.1250691.