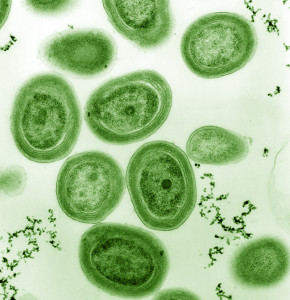
Single cell genomics reveals diversity of cyanobacteria subpopulations.
The Science:
Single cell genomics allowed researchers to examine the subpopulations of the marine microbe Prochlorococcus, affording a glimpse at “a new dimension of microdiversity.”
The Impact:
The cyanobacterial subpopulations were found to be distinct, comprised of “genomic backbones” made of highly conserved core gene alleles and a smaller set of flexible genes. The findings suggest that such genomic backbones may be a feature of diverse microbial populations.
Summary
Small but mighty, the tiny cyanobacterium Prochlorococcus is the foundation of the ocean food web. It is the smallest known photosynthetic organism and as such contributes significantly to the global carbon cycle. A recent study in the April 25, 2014 issue of Science suggests that the single species could be counted as consisting of thousands of species. A team led by Sallie Chisholm, a MIT researcher and longtime collaborator with the U.S. Department of Energy Joint Genome Institute (DOE JGI), applied single-cell genomics to study the cyanobacterial species. They sequenced and assembled Prochlorococcus genomes from single cells collected at the Bermuda-Atlantic Time-series Study site in the northwestern Sargasso Sea between November 2008 and April 2009.
“JGI is where the Prochlorococcus genomics story began about 15 years ago,” said Chisholm. “They sequenced the genomes of two of our strains that had been isolated from different oceans, giving us our first peek at the diversity within this group. Likewise, JGI’s current collection of a number of Prochlorococcus genomes has been an invaluable resource in our more recent work. These ‘reference genomes’ are key to unlocking the mysteries of the patterns we see in the genomes of wild populations.”
The team studied the subpopulations of Prochlorococcus at the single cell level and found that there were distinct “genomic backbones,” groups comprised of highly conserved core alleles, alternative forms of genes, and a smaller distinct set of flexible genes. Such a large set of coexisting subpopulations with distinct genomic backbones, the team concluded, may be characteristic feature of free-living bacterial species with huge populations in highly mixed habitats. The sequence variation found between the single cells sequenced and analyzed was verified with the help of DOE JGI’s Rex Malmstrom. “My contribution was to provide the essential control data,” he explained. “We could do this based on our extensive R&D work on single cell genomics.”
Contact
Sallie Chisholm
MIT
chisholm@mit.edu
Funding
- Department of Energy Office of Science
- National Science Foundation
- Gordon and Betty Moore Foundation Marine Microbiology Initiative
Publication
Kashtan N et al. Single-cell genomics reveals hundreds of coexisting subpopulations in wild Prochlorococcus. Science. 2014 Apr 25;344(6182):416-420. doi: 10.1126/science.1248575.