 Only three weeks ago the Department of Energy’s Joint Genome Institute announced completion of the working draft covering human chromosomes 5, 16 and 19. Today, JGI Deputy Director Trevor Hawkins was able to highlight a new milestone. In collaboration with Drs. Barbara Murray (UofT Houston Med. School) and George Weinstock (Baylor College of Medicine), the JGI had used its production sequencing facility to sequence the genome of “supergerm” Enterococcus faecium. Though the sequence data from this particular organism will doubtless be of great value to the scientific and medical community, even more noteworthy is the JGI’s ability to draft an entire microbe genome in a single day.
Only three weeks ago the Department of Energy’s Joint Genome Institute announced completion of the working draft covering human chromosomes 5, 16 and 19. Today, JGI Deputy Director Trevor Hawkins was able to highlight a new milestone. In collaboration with Drs. Barbara Murray (UofT Houston Med. School) and George Weinstock (Baylor College of Medicine), the JGI had used its production sequencing facility to sequence the genome of “supergerm” Enterococcus faecium. Though the sequence data from this particular organism will doubtless be of great value to the scientific and medical community, even more noteworthy is the JGI’s ability to draft an entire microbe genome in a single day.
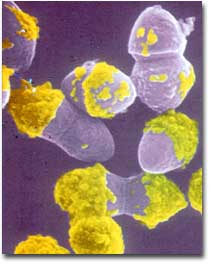
E. faecium is one of the top ten causes of nosocomial infections (infections obtained while hospitalized) in the United States. It can cause complicated abdominal infections, skin and skin structure infections, urinary tract infections and infections of the blood stream. In some cases infections can be life threatening, particularly in patients that are already immunodeficient. Previously found to be resistant to common antibiotics such as penicillin, strains were detected in the U.S. in 1989 that had also developed resistance to the “last resort” antibiotic vancomycin. Since this initial discovery, vancomycin-resistant enterococcus (VRE) has become a more and more prevalent concern, the number of cases increasing 20-fold between 1989 and 1993. The resistance to vancomycin is attributed to proteins, which are coded for in the genetic makeup of E. faecium. For this reason, knowledge of the complete genome sequence of this organism may be valuable in developing treatments for the infections it causes.