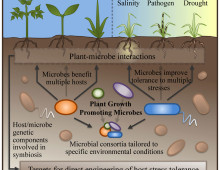
Researchers highlight the advantages of promoting beneficial plant-microbe relationships. The Science: The usual response to plant stressors such as low nutrient availability and disease has been to breed or develop cultivars that are tolerant or resistant to these concerns. In recent years, researchers have been paying more attention to the interactions between plants and microbial…
Dyeing to Learn More About Marine Viruses
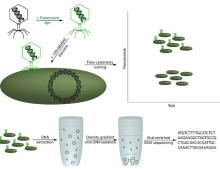
Tagging strategy allows for population surveys The sheer volume of cyanobacteria in the oceans makes them major players in the global carbon cycle and responsible for as much as a third of the carbon fixed. These photosynthetic microbes, which include Prochlorococcus and Synechococcus, are tiny – as many as 100 million cells can be found…
Soil microbiomes can set plant flowering time
Scientists study particular response to environment and its effect upon phenotype. The Science: Scientists grew Boechera stricta plants in soil inoculated with microbes from natural B. stricta habitats to study the flowering time phenotype. The Impact: The technique researchers employed to isolate soil microbes to study their effect on a single plant phenotype can potentially…
Treading into a Gray Area Along the Spectrum of Wood Decay Fungi
One of the most basic rules for playing the game “Twenty Questions” is that all of the questions must be definitively answered by either “yes” or “no.” The exchange of information allows the players to correctly guess the item in question. Fungal researchers have been using a variation of Twenty Questions to determine if wood-decaying…
Detecting evidence of selection
Genome-wide scan of maize population finds genes affected by long-term artificial selection. The Science: Researchers conducted a genome-wide scan of a long-term maize breeding study to find the genes involved in increasing the number of ears per maize plant. The Impact: The study demonstrates how significantly reduced costs associated with sequencing and the ability to…
IDing Livestock Gut Microbes Contributing to Greenhouse Gas Emissions
“Increased to levels unprecedented” is how the Intergovernmental Panel on Climate Change (IPCC) described the rise of carbon dioxide, methane and nitrous oxide emissions in their report on the physical science basis of climate change in 2013. According to the US Environmental Protection Agency (EPA), the atmospheric concentration of methane, a greenhouse gas some 28…
Eucalyptus—A Global Tree for Fuel and Fiber
From antiseptic oils to the construction of didgeridoos, the traditional Australian Aboriginal wind instrument, the eucalyptus tree serves myriad purposes, accounting for its status as one of the world’s most widely planted hardwood trees. Its prodigious growth habit has caught the eyes of researchers seeking to harness and improve upon Eucalyptus’ potential for enhancing sustainable…
When a stop sign is not interpreted as “stop”
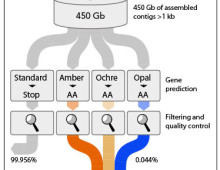
Microbes disprove long-held assumption that all organisms share a common vocabulary. The Science: Through single-cell genomics and metagenomics, researchers exploring the planet’s microbial diversity have found that not all organisms interpret a series of short genetic sequences to mean the same thing. The Impact: The ability to study microbes in the wild helps researchers realize…
More than just a hill of beans: Phaseolus genome lends insights into Nitrogen fixation
“It doesn’t take much to see that the problems of three little people doesn’t add up to a hill of beans in this crazy world,” Humphrey Bogart famously said in the movie Casablanca. For the farmers and breeders around the world growing the common bean, however, ensuring that there is an abundant supply of this…
Retracing early cultivation steps: Lessons from comparing citrus genomes
Citrus is the world’s most widely cultivated fruit crop. In the U.S. alone, the citrus crop was valued at over $3.1 billion in 2013. Originally domesticated in Southeast Asia thousands of years ago before spreading throughout Asia, Europe, and the Americas via trade, citrus is now under attack from citrus greening, an insidious emerging infectious…