The newest iteration of the DOE Joint Genome Institute’s and analytical tools sports improved user interface and infrastructure.
The Science:
The DOE Joint Genome Institute’s massive genomic database and data management system, the Genome Portal (http://genome.jgi.doe.gov), has recently been upgraded with a more robust infrastructure to manage the torrent of genomic data available and a variety of ways for users to access that information.
The Impact:
The amount of data now being generated by DOE JGI and its collaborators falls squarely in the big data category. The ramped-up supply and complexity of data means that the Genome Portal now needs a computational structure to keep pace. The details of those upgrades were outlined in an article in the January 1, 2014 issue of the journal, Nucleic Acids Research.
Summary
As one of the world’s top genome sequencing and analysis facilities – and the only one dedicated to energy and the environment – DOE JGI completed 2,635 projects in fiscal year 2012. That’s three times the number of projects they did in 2011. Their genome-sequence data consisted of more than 56 trillion nucleotides in 2012 and over 70 trillion nucleotides in 2013. DOE JGI’s clearinghouse for publicly available genomic data needed to be able to manage this output. In the last two years, DOE JGI has focused on improving the system for better data storage, access, downloads and analysis of the data.
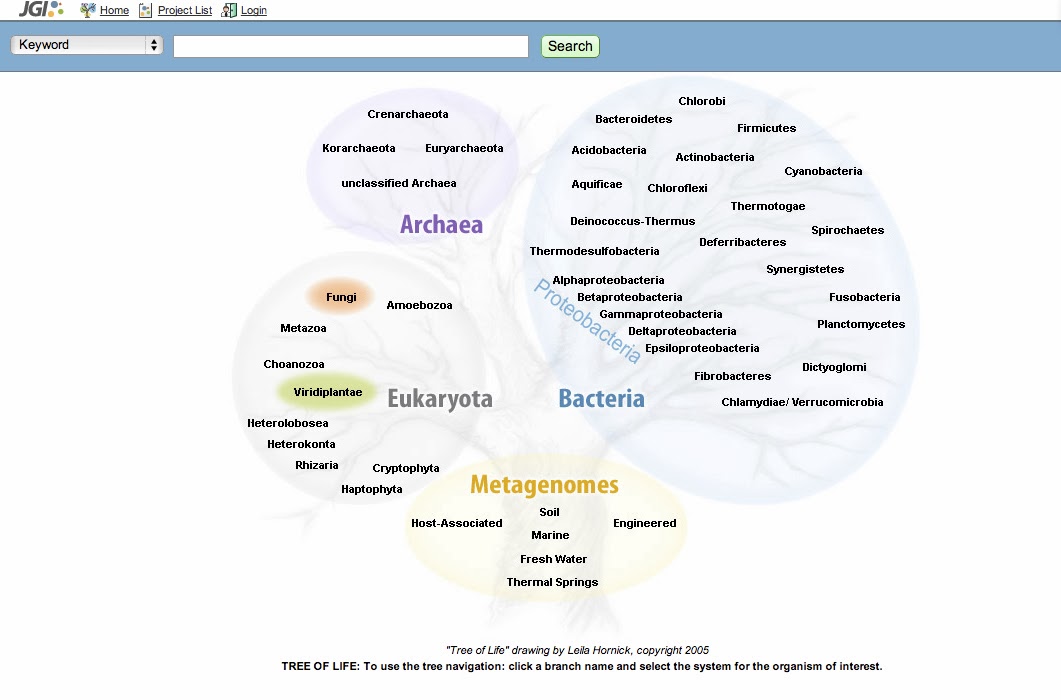
The new Genome Portal has several ways to access data, including a list of all DOE JGI projects, an improved search function and a “Tree of Life” visual search tool, which groups sequenced genomes by their domain of life and metagenomes by ecological niche. Users click on one of the “branches” and are redirected to the appropriate specialized database (IMG for microbial genomes, IMG/M for metagenomes, Mycocosm for fungal genomes and Phytozome for plant genomes). For genomes not belonging to those portals, users are routed to the individual portal pages for related projects.
The data in the Genome Portal is updated automatically as soon as it’s available through DOE JGI’s various annotation projects, cutting down on delays. DOE JGI’s higher volume of genome submissions led to a partnership between DOE JGI and NCBI which resulted in an automated submission process. Submissions to NCBI’s databases (Biosample,BioProject and GenBank) are now prepared automatically.
The Genome Portal now runs on servers hosted by the DOE’s National Energy Research Scientific Computing Center (NERSC) at Lawrence Berkeley National Laboratory. NERSC (http://www.nersc.gov/) is one of the nation’s foremost centers for high performance computing. JGI’s alliance with NERSC will mean faster and smoother access for users tapping into the Genome Portal’s resources.
Contact
Inna Dubchak
Joint Genome Institute
[email protected]
Publication
Henrik Nordberg, et al. The genome portal of the Department of Energy Joint Genome Institute: 2014 updates. Nucl. Acids Res. (1 January 2014)42 (D1): D26-D31 (published online ahead of print, November 12, 2013)
doi: 10.1093/nar/gkt1069
Funding
Department of Energy, Office of Science
Related Links
http://nar.oxfordjournals.org/content/42/D1/D26.long
The Genome Portal: http://genome.jgi.doe.gov