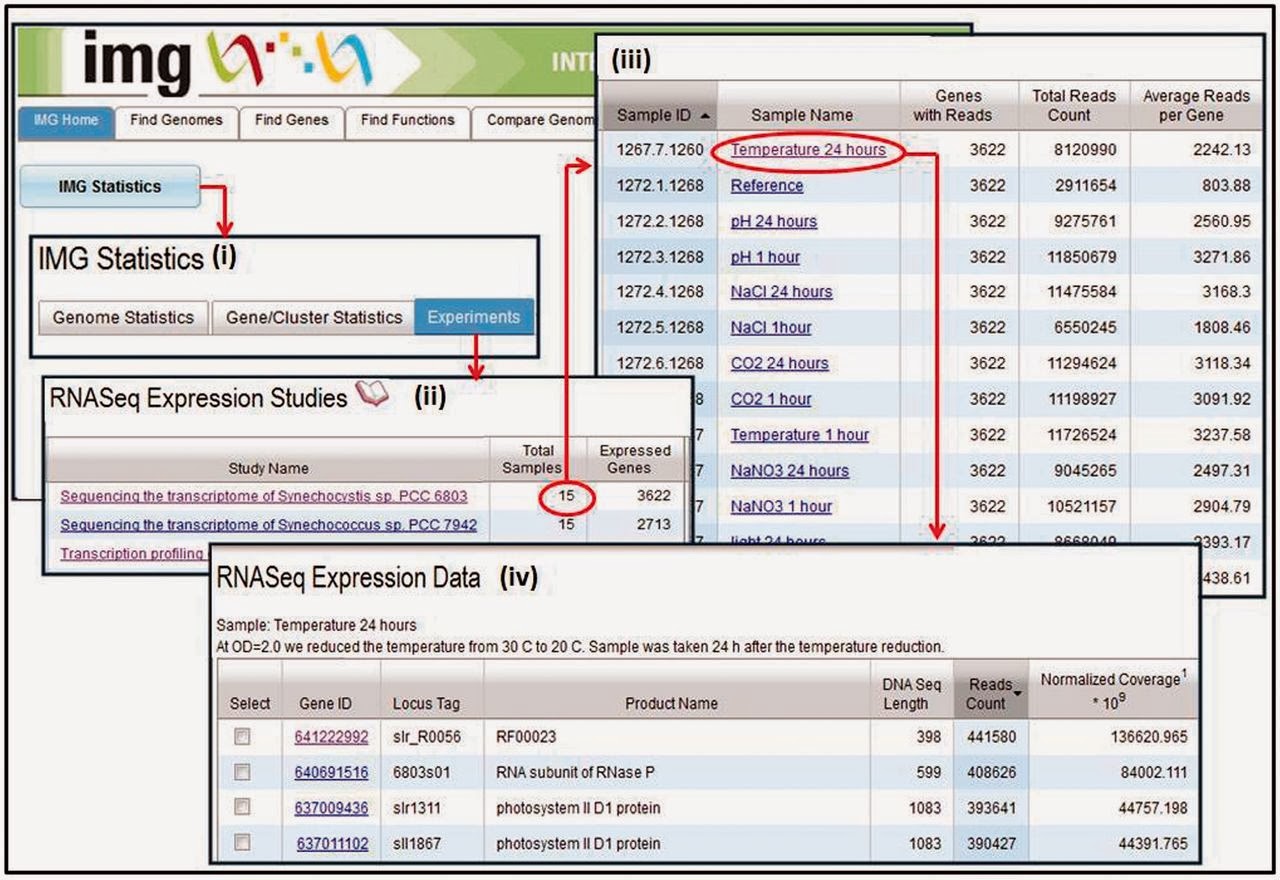
Capacities for DOE JGI’s twin genome analysis systems, IMG and IMG/M have both been expanded in the last two years.
The Science:
The DOE Joint Genome Institute maintains the Integrated Microbial Genomes (IMG) data warehouse, which contains a rich collection of genomes from all three domains of life. IMG/M provides a similar collection of microbial communities (metagenomes). Both have recently been upgraded to deal with the uptick in genome sequencing and provide more options for users.
The Impact:
One major challenge for users of IMG and IMG/M has been the swiftly growing number of genomes and metagenomes available for analysis. Both data systems have been cited in hundreds of publications and are also used by students learning genomics. Improvements in both systems have expanded their capacity and added new tools.
Summary
IMG was introduced almost a decade ago in 2005. Since the last published report in 2012, both systems have grown with the number of genomes and metagenomes and have been improved and refined as new tools have been roled out for the user community. The improvements for both systems were outlined in a pair of reports (http://nar.oxfordjournals.org/content/42/D1/D560.long and http://nar.oxfordjournals.org/content/42/D1/D568.long) in the January 1, 2014 issue of Nucleic Acids Research.
As of late 2013, the current version of IMG contains more than 16,000 genome datasets with more than 42 million protein-coding genes. Most (nearly 12,000) are bacterial, archaeal and eukaryotic genomes. That’s more than three times what the system housed two years ago. IMG also includes thousands of viral genomes, plasmids that did not come from a specific microbial genome sequencing project, and hundreds of genome fragments. At the same survey point, IMG/M contained 3,328 metagenome data sets from 460 metagenome studies, with >19.5 billion protein coding genes. About two-thirds of these metagenome data sets are publicly available to all users.
Both systems have enhanced analysis tools for publicly available datasets. The latest version of IMG includes tools for recording and analyzing single cell genomes, RNA sequencing data and gene clusters coding for synthesis of complex organic molecules (biosynthetic clusters).
Administrators for both systems are continually improving them to keep up with recent advances in genomics. Future advances include pangenome data (the full complement of genes that include core genes all individuals of a species have plus variant genes that help some individuals adapt to different environments) and analysis tools for IMG and metaproteomics datasets (protein samples collected from environmental sources) in IMG/M.
Contact
Nikos Kyrpides
Joint Genome Institute
[email protected]
Victor Markowitz
Joint Genome Institute
[email protected]
Publication
VM Markowitz et al. IMG 4 version of the integrated microbial genomes comparative analysis system. Nucl. Acids Res. (1 January 2014)42 (D1): D560-D567. First published online: October 27, 2013
doi: 10.1093/nar/gkt963
VM Markowitz et al. IMG/M 4 version of the integrated metagenome comparative analysis system Nucl. Acids Res. (1 January 2014)42 (D1): D568-D573. First published online: October 16, 2013
doi: 10.1093/nar/gkt919
Funding
Department of Energy, Office of Science
Related Links
IMG website: http://img.jgi.doe.gov/
IMG/M website: https://img.jgi.doe.gov/cgi-bin/m/main.cgi