Genome of world’s most common fungal symbiont sheds light on drought resistance role
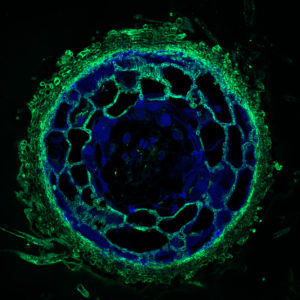
The crosscut shows the fungal tissues – the fungal mantle around the root tip and the fungal network of tendrils that penetrates the root of plants, or Hartig Net, between Pinus sylvestris plant root cells – in green. (Image by Maira de Freitas Pereira, INRA Nancy.)
The mutualistic relationship between tree roots and ectomycorrhizal (ECM) fungi has been shaping forest ecosystems since their inception. ECM fungi are key players supporting the growth, health and stress tolerance of forest trees globally, such as oak, pine, spruce, birch and beech, and help boost the productivity of bioenergy feedstock trees, including poplar and willow. The most common ECM fungus is Cenococcum geophilum, found in subtropical through arctic zones and especially in extreme environments. It is also the only mycorrhizal fungus in the Dothideomycetes, a large class comprised of some 19,000 fungal species, many of them plant pathogens.
To learn more about what ectomycorrhizal characteristics are dominant in Cenococcum geophilum, a team led by researchers at the French National Institute for Agricultural Research (INRA) and the Swiss Federal Institute for Forest, Snow and Landscape Research WSL, and including researchers at the U.S. Department of Energy Joint Genome Institute (DOE JGI), a DOE Office of Science User Facility, compared its genome with the genomes of close relatives Lepidopterella palustris and Glonium stellatum, neither of which are ECM fungi. The study was published online September 7, 2016 in Nature Communications. They found specific adaptations in the C. geophilum transcriptome – the set of its messenger RNA molecules that reflects actual biochemical activity by the fungus –that could help their hosts be more resistant to drought stress, a finding that could be useful in developing more plant feedstocks for bioenergy amidst the changing climate.
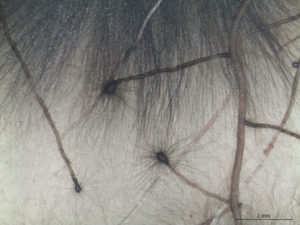
Shown here are poplar roots and the black mycelium and mycorrhizas formed by the fungus Cenococcum geophilum. (Martina Peter, Swiss Federal Research Institute WSL)
As part of a comparative genomic analysis done through the Mycorrhizal Genomics Initiative (MGI) headed by study senior author Francis Martin of INRA, the DOE JGI sequenced C. geophilum and its close relative Lepidopterella palustris, and annotated both of these genomes and another close relative, Glonium stellatum. “We showed that the genome of C. geophilum, the only known mycorrhizal symbiont within the largest fungal class Dothideomycetes, acquired the same genomic adaptations to the mycorrhizal lifestyle over generations as the previously sequenced ectomycorrhizal basidiomycetes,” Martin said. “These include a strikingly reduced number of plant cell wall degrading enzymes (PCWDEs) and a large set of symbiosis-induced lineage-specific genes, including dozen of mycorrhiza-induced small secreted effector-like proteins (MiSSPs).”Unlike free-living saprotrophs, fungi that get their nutrients from decomposing organic matter in forest soils and so require PCWDEs, Cenoccocum has come to rely heavily on its hosts for its carbon nutrition.
Key Genes for Drought Adaptation Found
Noting that the tree root tips colonized with C. geophilum are highly resistant to desiccation, one of the team’s key findings is that two of the three most highly induced C. geophilum genes in symbiosis code for water channels. “The regulation of these water channel genes is fine-tuned under drought conditions and they might therefore play a key role in drought adaptation of host plants,” said first author Martina Peter of the Swiss Federal Research Institute WSL.
“C. geophilum population genomics should shed light on the mechanisms of host and environmental adaptation,” the team wrote in their paper. “It should facilitate the identification of drought-adapted C. geophilum strains, which can be used to efficiently support their host trees threatened by the forecasted increase in drought periods in many parts of the world.”
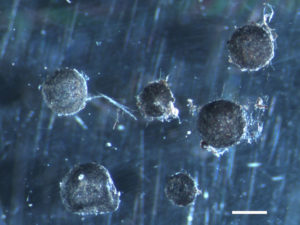
Shown here are the sclerotia, the resistant propagules formed and frequently found in soils. (Martina Peter, Swiss Federal Research Institute WSL)
Both Peter and Martin noted that credit for the success of the work and of the Mycorrhizal Genomics Initiative goes to the “tight collaboration” between their teams, the DOE JGI, Joey Spatafora at Oregon State University and Pedro Crous at the CBS Fungal Biodiversity Centre (Utrecht, Netherlands) as well as with Bernard Henrissat of CNRS and University of Aix-Marseilles.
“The intersection of genomics and evolutionary biology, as carried out in the MGI and the 1000 Fungal Genomes (KFG) Project, can inform our understanding of the biological principles intrinsic to mycorrhizal symbiosis,” said Martin. “By combining genome sequences with rigorous physiological and ecological studies, we are entering a time where linking the presence, composition and abundance of soil mycorrhizal communities with important soil processes and forest productivity at an ecosystem scale is possible.”