Microbes and their viruses are recently found to be profoundly impacting in virtually any ecosystem studied – from the oceans and soils to humans and bioreactors. While our knowledge of viral diversity is growing, we often observe genomes without knowing which host cell the virus infects. This proposal seeks to leverage ecosystem modeling and high-resolution time series datasets to scalably identify which viruses infect which hosts in experimental model systems and complex communities. Should it be successful, the new analytic will be powerful context for studying viruses in any ecosystem.
Microbial Genomes Across the World’s Rivers
This proposal seeks to create a global census of microbial genomes across 250 of the world’s rivers, from the Amazon to the Mississippi. Our goal is to create a Genome Resolved Open Watershed (GROW) database- which will be an open access resource to advance knowledge of aquatic microbiology for the entire scientific community.
Inter-organismal Interactions in the Rumen Ecosystem
We propose to generate data that would provide us with a better understanding of the role of different microorganisms involved methanogenesis in the rumen ecosystem and the foundation to develop new strategies for methane mitigation from ruminants. Ruminant animals are one of major anthropogenic sources of the highly potent greenhouse gas methane and advanced methane mitigation strategies would have a significant impact on the global methane emission and the therefore on climate change. This aligns fully with DOE’s mission to reduce the anthropogenic carbon footprint.
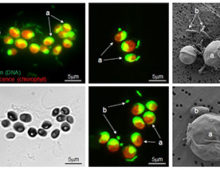
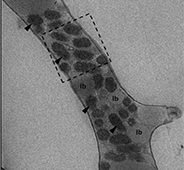
Diversity of Parasitic and Commensal Chytrids
Chytrids are basal fungi. Despite the chytrids major negative impact on algal phytoplankton productivity and as a major pest in microalgal biofuels production facilities, extremely little is known about the biology of these organisms. We expect that the genomic data provided from this CSP will shed light on the “blackbox” of unknown molecular events at various stages of infections in the inter-organismal chytrid/algal systems. This work is highly relevant to the DOE mission in the context of organic carbon cycling in aquatic plankton communities and due to the microalgae being feedstock for biofuels applications.
International Arctic Ice Drift Experiment MOSAiC
The MOSAiC drift experiment will provide sequence data to study for the first time how microbial communities change over a complete seasonal cycle in the Arctic Ocean in terms of their diversity and gene activity. These results will be instrumental to study climate processes they drive.
Cross-Domain Interactions in Microalgae Communities
We will characterize algal communities from some of the most productive ecosystems in the world to understand the interactions between the microbial community members involved. Our goal is to develop strategies for establishing stable and highly productive photosynthetic communities suitable for biofuel and high-value product generation on an industrial scale.
Characterizing Viruses in Antarctica
Characterizing viruses in extreme environments may facilitate a better understanding of the role that viruses play in the dynamics and evolution of complex microbial communities.
Role of Fungal Endosymbionts in Host Reproduction
Some beneficial plant associated fungi house bacteria inside their cells which impact fungal and plant health. Recent studies show that some fungal endosymbionts manipulate reproduction of their hosts, and may influence plant ecosystems. This project focuses on endosymbionts and fungal reproduction, expanding knowledge on the complexities of microbiome interactions. These fungi are industrially important because they produce valuable lipid products. Our goal is to learn about optimal conditions for fungal health and apply that knowledge to increase crop health and produce alternative energies.
Genomic Variation in Mustard Family Metabolism
Studies of genomic variation are critical to developing a synthetic, systems and quantitative understanding of how to improve crops. However, modern short-read methodologies may actually miss the very genes that are critical to adapting crops to new environments or making them healthier. This project will create high-quality genomes of multiple crops and species from the mustard family to test new methods and technologies to ensure that these genes essential for adaptation and improvement are identified.
Expanding Metabolic Understanding of C- and S- Cycling Microbes
Is there more biochemistry to be discovered in the microbial world? How universal is biochemistry between distantly related bacteria? A massive amount of microbial genome data exists, yet every new sequence finds genes for which we cannot predict a concrete function. This proposal seeks to combine genomic information with direct detection of metabolites to answer these questions and provide a path for assigning gene functions in two distantly related microbial groups that play key roles in carbon and sulfur cycling in multiple ecosystems across the planet.